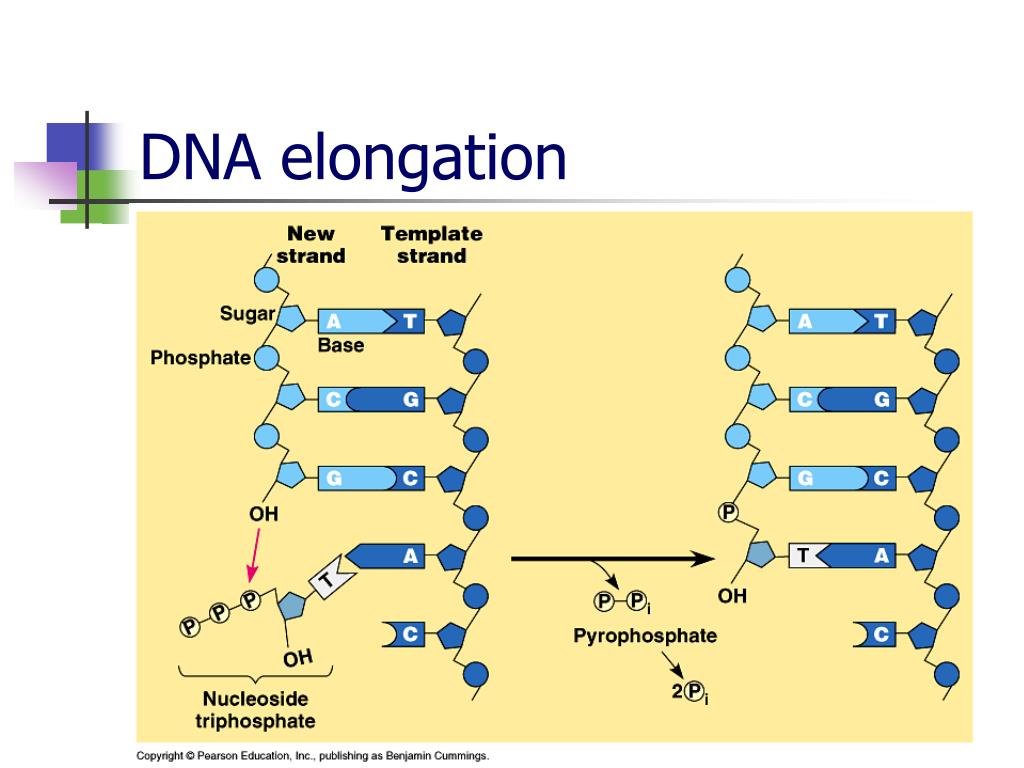
  For bidirectional replication, thisrequires only one origin, and indeed this is the case. Dividing the size of the chromosome by this amountsynthesized per fork gives 4.64 ´ 10 6bp / 2.4 ´ 10 6 bp, or 1.93.Hence two replication forks are sufficient. Thus in 40 min,one replication fork replicates 60,000 bp per min ´ 40 min = 2.4 ♁0 6 bp. As the replication fork moves 60,000 nucleotidesper min, it produces both daughter strands at the same rate. The reverse of the polymerizationreaction is pyrophosphorolysis (adding a molecule of pyrophosphate across thebond that is broken), resulting in a nucleoside tri phosphate and a DNA chain shorter by one nucleotide.Īnswer 5.6. Removal of a nucleotide from the 3’ end of thegrowing chain by a 3’ to 5’ exonuclease catalyzes the hydrolysis of the 3’nucleotide (adding a molecule of water across the bond that is broken),generating a nucleoside mono phosphateand a DNA chain shorter by one nucleotide. 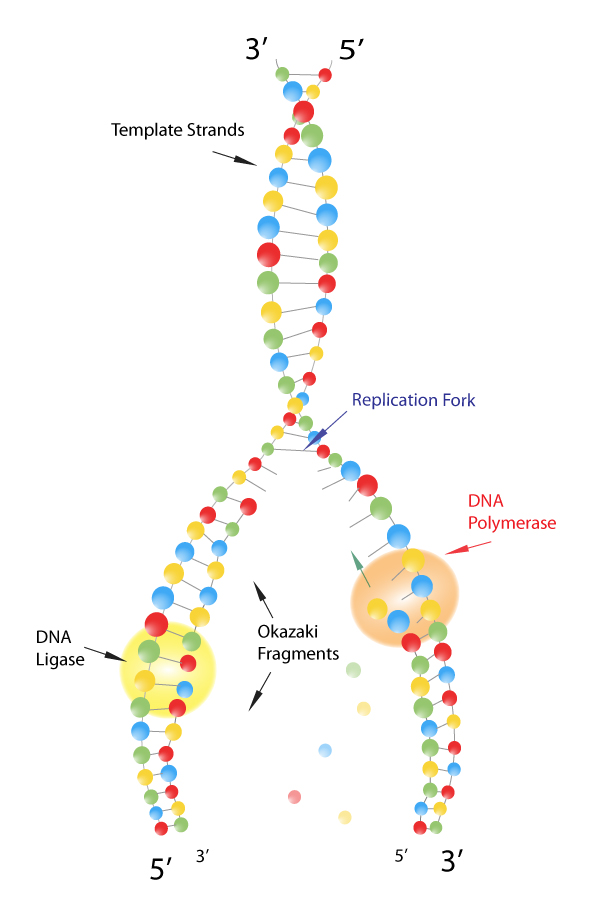
Note thatif an incorrectly incorporated nucleotide were removed by a proofreadingexonuclease (a 5’ to 3’ exonuclease in this hypothetical example), then theactivated end of the chain would be removed, and synthesis would stop.Īnswer 5.5. Chain synthesis would occur in a 3’ to 5’ direction. All these steps are similar to those inthe tail-growth mechanism at the 3’ end, except that the nonactivated end ofthe incoming nucleotide initiates the reaction with the activated end of thegrowing chain. The b- and g-phosphateswould be liberated as pyrophosphate. The 3’ hydroxylon an incoming nucleotide could react with the a-phosphate of the 5’ nucleotide by a nucleophilic attack. 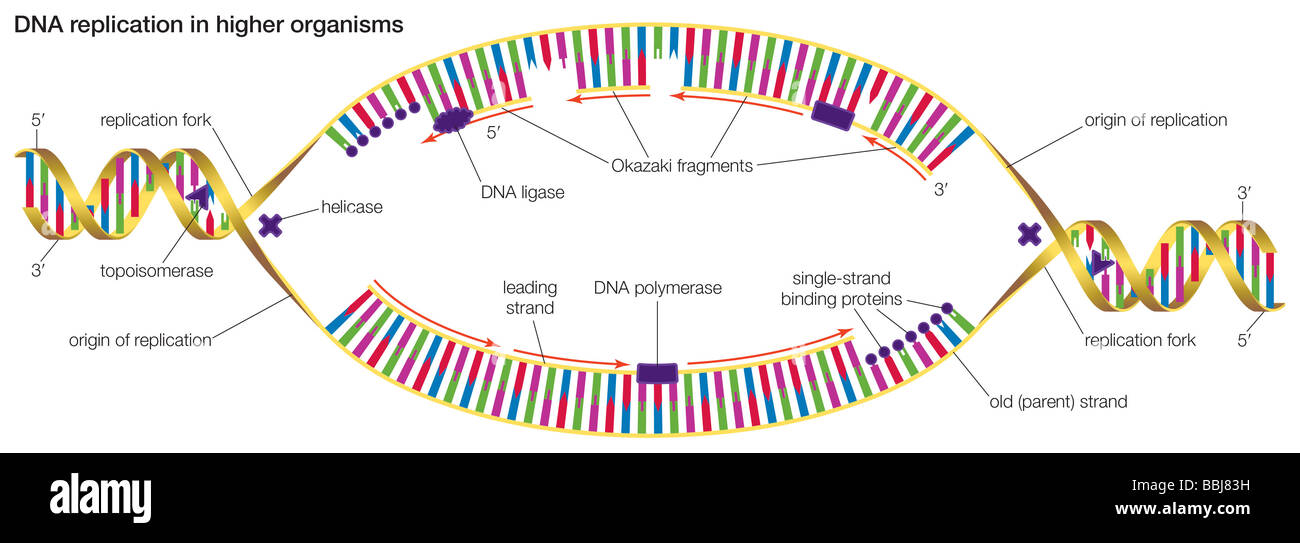
A hypothetical head-growth mechanismfor DNA synthesis would have the 5’ end of the primer at the active site this5’ end would have a triphosphate on the last nucletide added. 5.7 you would not expect to see a slow-sedimenting peakof nascent DNA.Īnswer5.4. Therefore, by the model of semidiscontinuoussynthesis shown in Fig. Because the short Okazaki fragments shouldstill be in duplex with the large parental DNA strands, the duplex would notseparate from the bulk of the DNA. In an neutral sucrose gradient, the two strands ofthe DNA duplex should stay together. In contrast to the replication eyes, the two new strands are notsynthesized simultaneously at the replication fork in D loop replication.Īnswer 5.3. Theproduction of LL shows that replication is not random.Īnswer5.2. Une série d'expériences conduit à la découverte des fragments d'Okazaki.Answer 5.1. Okazaki fragments meaning series#L'invention concerne un procédé pour isoler la séquence d'ADN codant pour la L-asparaginase recombinante de la bactérie Erwinia carotovora.Ī series of experiments led to the discovery of Okazaki fragments. The invention relates to the secretion of the Okazaki fragment which codes L-asparaginase of the Erwinia caratovora bacterium. Lorsque l' ADN polymérase atteint l' amorce du fragment d' Okazaki suivant, l'amorce ARN synthétisée par la primase est hydrolysée et remplacée par de l'ADN. Les clamps peuvent également se lier à d'autres facteurs intervenant dans l'homéostase génomique, tels que les facteurs d'assemblage des nucléosomes, les ADN ligases des fragments d'Okazaki et les protéines de réparation de l'ADN.Įach Okazaki fragment is preceded by an RNA primer, which is displaced by the procession of the next Okazaki fragment during synthesis. Okazaki fragment Okazaki fragments are short, newly synthesized DNA fragments that are formed on the lagging template strand during DNA replication.įragment d'Okazaki Les fragments d'Okazaki sont des fragments d'ADN courts et nouvellement synthétisés qui sont formés sur le faisceau de duplication durant la réplication de l'ADN.ĭNA clamps also associate with other factors involved in DNA and genome homeostasis, such as nucleosome assembly factors, Okazaki fragment ligases, and DNA repair proteins. Finally, each new Okazaki fragment is attached to the completed portion of the lagging strand in a reaction catalyzed by DNA ligase.Įn conclusion, chaque éclat neuf d' Okazaki est fixé à la partie réalisée de la boucle de ralentissement dans une réaction catalysée par la ligase d'ADN. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
